I can’t find anything online about the results of the regional Breadfruit Biocultural Conservation Knowledge Exchange Workshop organized by the Pacific Island Farmers Organisation Network (PFO) at the Tutu Rural Training Centre in Fiji from 27–29 April 2026. Beyond social media posts, that is. But I really like the idea of participants sharing the breadfruit varieties that are special to them.
Stairway to maize diversity
There’s a nice article in Rising Kashmir highlighting that region’s cold-tolerant maize landraces as a unique source of genetic diversity. What I liked about it is that it doesn’t condescend to its audience. It’s unapologetically technical and niche, while successfully (I think) striving to be understood by all. That’s rare. The author, Dr Salika Ramazan, argues that long adaptation to Himalayan environments has produced valuable traits for climate resilience and future maize breeding, and advocates for urgent conservation before this irreplaceable diversity is lost.
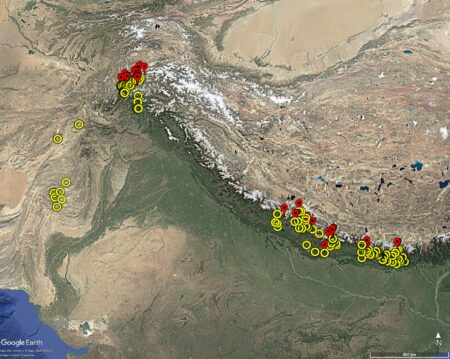
A quick search on Genesys revealed 302 maize accessions from above 1500 masl in the Himalayas (yellow on the map below), and 62 above 2500 masl (red). Of course, there are many more maize accessions from high altitudes in Central and South America, but their photoperiod adaptation (among other things) is likely to be quite different.
Brainfood: Markets edition
- “What is the essence of cultivating a crop that does not yield enough to feed my family?” Farmer agency and the management of agrobiodiversity in Ghana and Burkina Faso. Farmers balance subsistence needs and market opportunities when deciding which crop varieties to maintain. Well I never.
- Reassessing the economic returns of diverse traditional agricultural systems for smallholder farmers: a case study of the milpa in Mexico. Diverse traditional farming systems can generate significant economic value; so no, agricultural diversity is not necessarily less profitable than specialization for market production.
- Farmer participatory evaluation of Amaranthus cruentus L. breeding lines for marketable vegetable yield and organoleptic quality under on-farm and on-station conditions. In any case, farmers can work with researchers to select amaranth varieties with traits that improve both marketability and consumer appeal, linking crop improvement directly to market demand.
- Novel soybean type with improved volatile and sensory characteristics of raw soy slurries. Breeding soybean varieties with enhanced sensory qualities can hopefully increase consumer acceptance and create new market opportunities for soy-based foods. Unclear if farmers were involved, but they could have been..
- Can the digital long tail effect in farmers’ markets increase crop diversity on farms and in diets? Yes. Digital platforms can connect niche producers and consumers, creating markets for a wider range of crops, thereby encouraging agricultural and dietary diversity. How about linking seed producers to farmers?
- Preserving crop genetic diversity through traditional seed systems: insights from farmer-saved fonio (Digitaria exilis) landraces in Northern Ghana. Farmer-managed seed systems support the conservation of crop diversity while maintaining access to locally adapted varieties with potential market value. But maybe they could use a digital platform?
- Seeds and social norms: sorghum seed exchange among smallholder farmers in Northern Ethiopia. Cultural practices shape how farmers share seeds, influencing the circulation of crop diversity and farmers’ participation in local seed markets. Good luck with those digital platforms.
Brainfood: Indigenous edition
- Rapid adaptive increase of amylase gene copy number in Indigenous Andeans. Indigenous Andean populations evolved exceptionally high copy numbers of the AMY1 salivary amylase gene, likely linked to long-term adaptation to starch-rich diets associated with potato domestication roughly 10,000 years ago.
- Horse genetics, archaeology, and the beginning of riding. Horse domestication was not a sudden genetic event beginning around 2200–2100 BCE, but a long and regionally varied process in which Indigenous Eurasian pastoralists progressively managed, rode, milked and selectively bred multiple horse lineages over many centuries, transforming mobility and social organization well before the rise of the dominant modern domestic horse lineage.
- Bridging biodiversity and food systems: A nationwide synthesis of non-conventional food plants (PANCs) in Brazil. Brazil’s non-conventional food plants (PANCs) and associated Indigenous and traditional knowledge could help build more diverse, climate-resilient and socially inclusive food systems while strengthening biodiversity conservation, rural livelihoods and public food programs.
- Indigenous Wisdom for a Changing World: Bridging Traditional Ecological Knowledge and Biodiversity Conservation. Sacred groves and other community-managed landscapes in central Ethiopia conserve high levels of biodiversity through Indigenous institutions, ritual practices and traditional ecological knowledge, suggesting that effective conservation depends on treating cultural stewardship systems as integral to ecological resilience rather than as secondary to scientific management.
- When Knowledge Isn’t Free: Legal and Ethical Imperatives of Protecting Indigenous Intellectual Property. There’s a persistent mismatch between Western intellectual-property regimes and Indigenous concepts of collective ownership, biocultural heritage and intergenerational custodianship of knowledge, and that’s unfair.
- Crediting and citing Indigenous Knowledges within research. Biodiversity conservation becomes more effective when Indigenous scientists and communities participate as equal partners rather than merely as local stakeholders or informants.
MLS spoiler alert
Remember that “A comprehensive analysis of the operations of the Multilateral System – Insights and key figures 2025” that we trailed a few months back? The official launch is on 22 May and you can follow along virtually.
LATER: And these are the key takeaways, according to the Plant Treaty secretariat…

